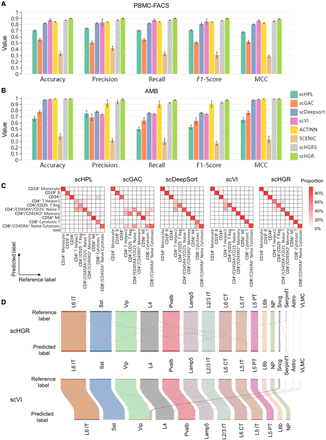
Performance comparison of scHGR and state-of-the-art methods on intra–data sets. Involving four state-of-the-art techniques, the experiments are conducted separately on PBMC-FACS (A) of human and AMB (B) of mouse and take the mean value of fivefold cross-validation to generate the histogram. Each group represents an evaluation metric, and each color indicates a method. The segment at the top of each bar represents the error segment of the corresponding method. (C) Heatmap of the confusion matrix of PBMC-FACS, with rows denoting predicted labels by the corresponding method and columns denoting labels provided by reference data sets. Notably, scHPL assigns some cells as root results from its tree architecture. (D) For AMB, the Sankey diagram compares cell type assignments between reference data sets and predicted labels from scHGR and scVI.











