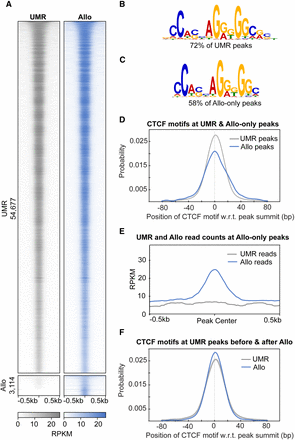
Allo results in the discovery of 3114 additional peaks in a CTCF ChIP-seq data set. (A) ChIP-seq heatmaps comparing K562 CTCF peaks called by MACS2 using UMRs only or UMRs plus Allo-mapped reads. (B,C) Top-ranked motifs from MEME-ChIP in the UMR-derived and Allo-only peaks, respectively. (D) Position of the CTCF motif with respect to the peak summit for UMR-derived peaks and Allo-only peaks. (E) Read depth at Allo-only peaks using UMR BAM files versus using Allo output. (F) Position of the CTCF motif with respect to the peak summit before and after the inclusion of Allo at UMR-derived peaks.











