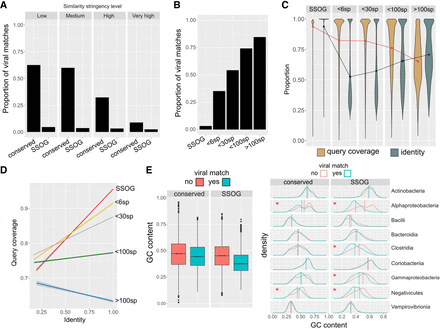
Properties of genes with viral matches. (A) Proportions of conserved proteins and SSOGs with statistically significant similarity matches to viral proteins at four different significance thresholds. (B) Proportions of proteins with statistically significant similarity matches to viral proteins (at high stringency level), binned according to the number of prokaryotic species (sp.) in which a homolog can been found (size of the prokaryotic protein family). (C) Query (prokaryotic protein) coverage and sequence identity of the top statistically significant matches (all hits with E-value < 10−5 are included), binned according to the number of species in which a homolog can be found. Lines connect distribution means. (D) Correlation between query coverage and identity in sequence matches involving genes belonging to different sizes of protein families, same data as in C. (E) Distributions of GC content of genes with and without a statistically significant viral match (high stringency), among all SSOGs and conserved genes (boxplot) and among the nine best represented taxonomic classes (density plots). A red asterisk denotes a nonnegligible effect size in difference of means calculated by Cliff's Delta (Delta estimate > 0.15). Note that in Alphaproteobacteria and Negativicutes, the difference exists also in conserved, but the effect size there is much weaker compared with SSOGs (−0.21 vs. −0.62 and −0.2 vs. −0.42).











