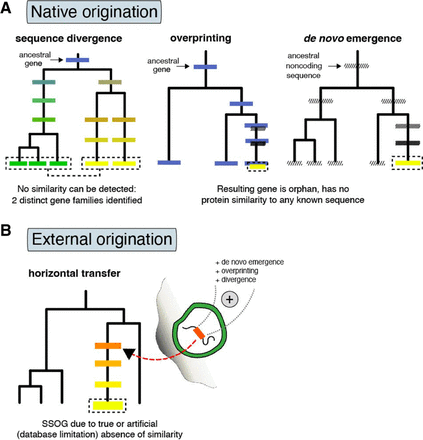
Evolutionary routes to SSOGs in prokaryotes. (A) Native origination through sequence divergence, as well as de novo emergence from entirely noncoding regions (known as de novo emergence) or from coding regions in alternative reading frames (known as overprinting). Overprinting can also be succeeded by gene duplication, such that both genes are then encoded in different regions. Darker shading represents transition from noncoding state to a coding one. Different colors represent differences in sequence similarity between homologous genes or in the same gene over time. (B) Origination in phages, plasmids, or integrative elements by various mechanisms such as de novo birth, overprinting, or divergence, and subsequent transfer to prokaryotic genomes. Because of either technical limitations or rapid divergence, similarity to the source of such externally originated genes might be lost, which would result in them being perceived as species-specific. Additionally, remodeling of existing genes, often in combination with overprinting and sequence divergence, can lead to SSOGs (not shown here).











