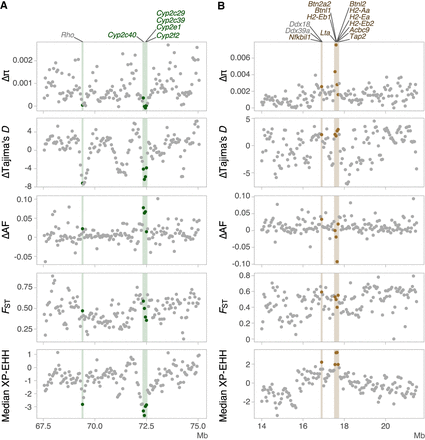
Examples of genomic landscapes around the regions for which hard selective sweeps were detected in common voles from Orkney or from the European continent. Δπ (πcontinent – πOrkney), ΔTajima's D (Tajima's Dcontinent –Tajima's DOrkney), ΔAF (AFcontinent – AFOrkney), FST, and median XP-EHH are shown. Each point represents one 50 kb window. Positively selected windows were marked with vertical lines. Genes overlapping with positively selected regions (outlier windows determined with DBSCAN with the highest or lowest one percentile of median XP-EHH value) are given on top. (A) Positively selected regions of continental voles had low Δπ, ΔTajima's D, and ΔAF. Genes belonging to the epoxygenase P450 pathway on M. arvalis Chromosome 4 are marked with green. (B) In Orkney voles, Δπ, ΔTajima's D, and ΔAF of positively selected regions did not form clusters outstanding from the genomic background. Genes related to antigen processing and presentation on Chromosome 15 are marked with brown.











