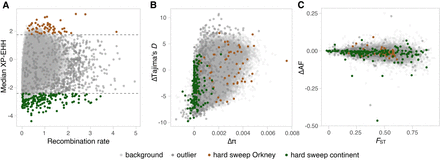
The distributions of genomic parameters used for tests of positive selection with DBSCAN. Each dot refers to one 50 kb window. (A) The median XP-EHH value of the outlier windows, which measures relative haplotype homozygosity, was negatively correlated with recombination rate in Orkney voles (Pearson coefficient r = −0.33) and positively correlated with recombination rate in continental voles (Pearson coefficient r = 0.38). The dashed lines show the threshold of the highest and lowest 1% of XP-EHH. (B) Regions positively selected only in continental voles had lower Δπ and ΔTajima's D, whereas regions positively selected only in Orkney showed large variation. (C) The major allele frequency difference of the regions under a hard selective sweep showed, overall, no obvious deviation from the genomic background (K–S test, P = 0.32, Z-test P = 0.19), but mean FST values were lower than the average over the genome (Z-test P = 0.0001).











