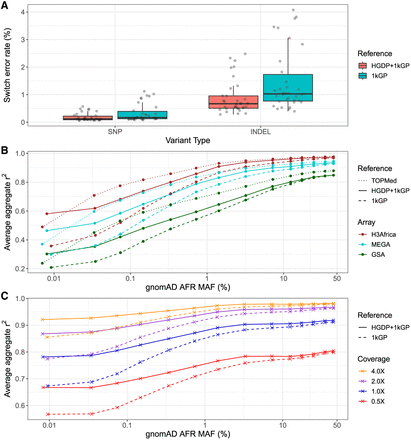
Phasing and imputation accuracy are improved across data generation strategies compared with existing reference panels. (A) Switch error rates for SNPs and indels in a truth set of 34 HGSVC2 genomes when using HGDP + 1kGP versus 1kGP reference panels for phasing. (B,C) Imputation performance as a function of minor allele frequency (MAF) for AFR in gnomAD v3.1 data using TOPMed, HGDP + 1kGP, and 1kGP reference panels in SNP array (B) and low-coverage sequencing data (C). Aggregate r2, which is the correlation between the imputed dosages and high-coverage “truth” genotype calls, was computed in MAF bins and averaged across Chromosomes 1–22. The validation set is composed of 93 AFR individuals sequenced at 30× coverage (Martin et al. 2021).











