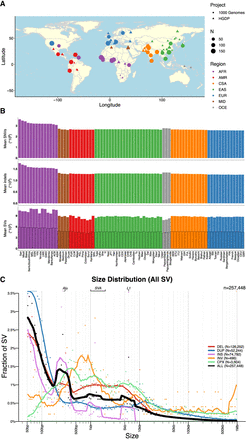
Geographical locations and genetic variants across populations. (A) Global map indicating approximate geographical locations where samples were collected. Coordinates were included for each population originating from the Geography of Genetic Variants browser as well as metadata from the HGDP (Marcus and Novembre 2017; Bergström et al. 2020). (B) Mean number of SNVs (top panel), indels (middle panel), and SVs (bottom panel) per individual within each population. The bottom panel showing SVs has a set of outlined bars in black that counts the subset of SVs called outside of highly repetitive genomic regions, which decreases calling accuracy in short-read sequencing data (Zhao et al. 2021). The bars without the outline indicate total SV counts regardless of whether they span repetitive regions. (A,B) Colors are consistent with geographical/genetic regions as follows: (AFR) African, (AMR) admixed American, (CSA) Central/South Asian, (EAS) East Asian, (EUR) European, (MID) Middle Eastern, and (OCE) Oceanian. (C) Sizes of SVs decay in frequency with increasing size overall with notable exceptions of mobile elements, including Alu, SVA, and LINE-1. (DEL) Deletion, (DUP) duplication, (CNV) copy number variant, (INS) insertion, (INV) inversion, and (CPX) complex rearrangement.











