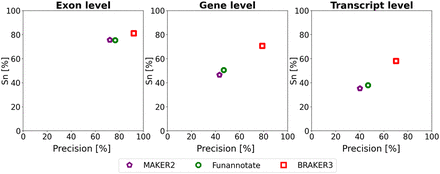
Average precision and sensitivity of gene predictions made by MAKER2, Funannotate, and BRAKER3 for a subset of eight species (excluding the mouse, spider, and fish genomes). Inputs were the genomic sequences, short-read RNA-seq libraries, and protein databases (close relatives included). The accuracy of MAKER2 reported here can be regarded as an upper limit of what can be expected when annotating a previously unannotated genome (see “Experiments” section).











