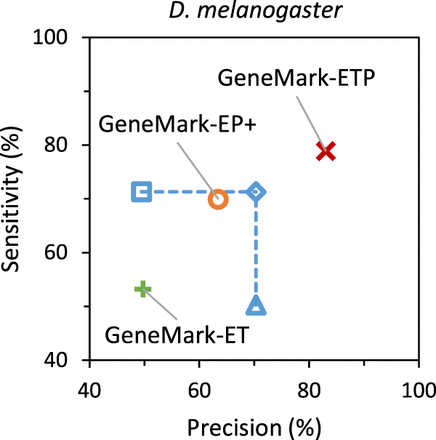
Gene-level Sn and Pr of the gene predictions in the D. melanogaster genome. The gene predictions were generated by GeneMark-ET (green), GeneMark-EP+ (orange), and GeneMark-ETP (red). We also show the Sn and Pr values for the Union (square) and the Intersection (triangle) of the sets of GeneMark-ET and GeneMark-EP+ gene predictions and the “best combination” of these two gene-prediction sets (diamond). The true positives were defined with respect to the annotation of the D. melanogaster genome. The “species excluded” database was used for the GeneMark-EP+ and GeneMark-ETP runs.











