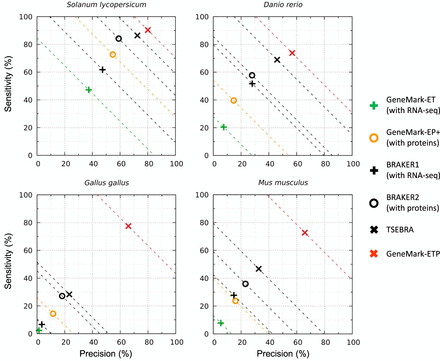
Gene-level Sn and Pr of gene predictions made by GeneMark-ETP and five other gene finders in the four large genomes: S. lycopersicum, D. rerio, G. gallus, and M. musculus. The Sn and Pr values for each genome were defined with respect to subsets of genes present in the RefSeq and Ensembl genome annotations (the intersections and the unions of annotations; see text). For these genomes, gene candidates lacking extrinsic support did not make it into the GeneMark-ETP list of predicted genes (see text). The “order excluded” protein databases were used.











