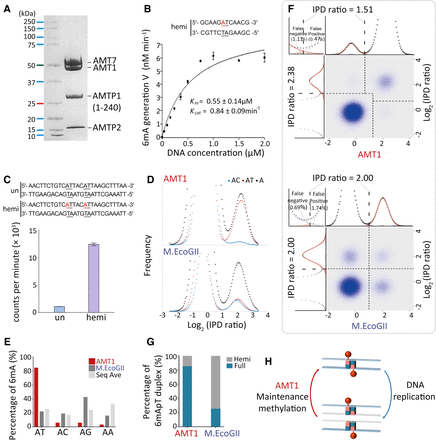
In vitro MTase activity of AMT1 complex. (A) SDS-PAGE of in vitro reconstituted AMT1 complex comprising AMT1, AMT7, AMTP1 (1–240 aa), and AMTP2. (B) The steady-state kinetics of AMT1 complex on a hemi-methylated substrate (hemi), determined by a 3H-SAM-based MTase assay. The substrate contains a single ApT duplex (underlined), which is hemi-methylated (red). (C) Methylation of the unmodified (un) and hemi-methylated (hemi) substrates. Both contain two ApT duplexes (underlined), which are either unmodified or hemi-methylated (red). (D) IPDr distributions for total adenine, adenine at the ApT dinucleotide, and adenine in ApC dinucleotide, after in vitro methylation of human chromatin by either AMT1 complex (top) or M.EcoGII (bottom). (E) 6mA distribution at all four ApN dinucleotides, after in vitro methylation of human chromatin by either AMT1 complex or M.EcoGII. ApN frequencies in SMRT CCS read are also plotted for comparison (Sequence Average). (F) Demarcation of the four methylation states of ApT duplexes by their IPDr on W and C, in human chromatin methylated by AMT1 complex (top) or M.EcoGII (bottom). AMT1 complex methylation pattern is reminiscent of that in WT Tetrahymena cells, with a strong preference for full-6mApT, as indicated by a shift in the IPDr threshold for calling full-6mApT relative to calling bulk 6mA. M.EcoGII methylation pattern is reminiscent of that in ΔAMT1 cells, with no preference for full-6mApT, as indicated by the same IPDr threshold for calling bulk 6mA or full-6mApT. (G) Relative abundance of hemi-6mApT and full-6mApT in human chromatin methylated by either AMT1 complex or M.EcoGII. (H) Model: AMT1-dependent semiconservative transmission of 6mA.











