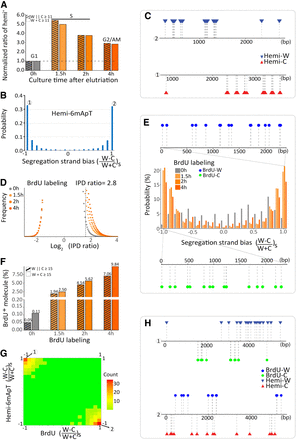
Segregation of hemi-6mApT to the parental strand after DNA replication. (A) Hemi+ molecules are enriched in S phase. Tetrahymena cells were synchronized at G1 phase by centrifugal elutriation and released for growth in the fresh medium (Liu et al. 2021b). Four time points were taken (0, 1.5, 2, and 4 h after release) for SMRT CCS. Hemi+ molecules were defined as DNA molecules with a total count of no less than 11 hemi sites (W + C ≥ 11) or with no less than 11 hemi sites on one strand (W||C ≥ 11). The count of hemi+ molecules was normalized first against the counts of total DNA molecules and then against the 0 h (G1 phase) value. (B) Hemi-6mApT sites in hemi+ molecules exhibit strong segregation strand bias. Segregation strand bias for hemi-6mApT is defined as the difference-sum ratio between hemi-W and hemi-C: [(W − C)/(W + C)]s. (C) Typical DNA molecules with hemi-6mApT fully segregated to W or C, corresponding to segregation strand bias of +1 and −1, as marked in B. (D) IPDr distributions of T sites in genomic DNA samples of synchronized Tetrahymena cells with BrdU labeling (1.5 h, 2 h, and 4 h) or without (0 h). The IPDr threshold for calling BrdU was set at 2.8. (E) BrdU sites in BrdU+ molecules exhibit strong segregation strand biases. BrdU+ molecules were defined as DNA molecules with a total count of no less than 15 BrdU sites (W + C ≥ 15). Segregation strand bias for BrdU was defined as the difference-sum ratio between BrdU sites on W and C: [(W − C)/(W + C)]s. Also shown are typical BrdU+ molecules with BrdU fully segregated to W or C, corresponding to segregation strand bias of +1 and −1, respectively. (F) Correlation between BrdU labeling and BrdU+ molecules. BrdU+ molecules were alternatively defined as DNA molecules with a total count of no less than 15 BrdU sites (W + C ≥ 15), or with no less than 15 BrdU sites on one strand (W||C ≥ 15). The latter is more selective for DNA molecules with strong strand segregation bias. (G) Hemi-6mApT and BrdU are segregated to opposite strands of the DNA duplex. Distribution of hemi+/BrdU+ molecules (hemi-6mApT: W||C ≥ 11; BrdU: W||C ≥ 15) according to their segregation strand bias for hemi-6mApT and BrdU, respectively. (H) Typical hemi+/BrdU+ molecules with hemi-6mApT and BrdU fully segregated to opposite strands, corresponding to segregation strand bias of (−1, +1) and (+1, −1), as marked in G.











