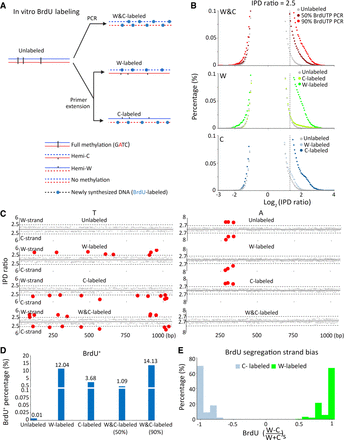
SMRT CCS detection of BrdU incorporation into newly synthesized DNA. (A) In vitro BrdU labeling. Specific labeling of either strand (W-labeled or C-labeled) was achieved by primer extension, whereas labeling of both strands (W&C-labeled) was achieved by PCR. A plasmid fragment containing three fully methylated GATC sites (6mA) was used as the template, as well as the unlabeled control for SMRT CCS. (B) IPDr distributions (log2) of all T sites from: both strands in unlabeled and W&C-labeled DNA (50% or 90% BrdUTP; top); only W in unlabeled, W-labeled, and C-labeled DNA (middle); only C in unlabeled, W-labeled, and C-labeled DNA (bottom). IPDr threshold was set at 2.5 for separating BrdU from T. (C) IPDr for all T (left) or A sites (right) in typical SMRT CCS reads for unlabeled, W-labeled, C-labeled, and W&C-labeled DNA (90% BrdUTP). IPDr thresholds were set at 2.5 for separating BrdU from T, and at 2.7 for separating 6mA from A. (D) Percentage of BrdU+ molecules in unlabeled, W-labeled, C-labeled, and W&C-labeled DNA (50% and 90% BrdUTP, respectively). BrdU+ molecules were defined as DNA molecules with no less than eight BrdU sites on one strand (W||C ≥ 8). (E) Segregation strand biases of BrdU sites in BrdU+ molecules. Segregation strand bias for BrdU was defined as the difference-sum ratio between BrdU sites on W and C: [(W − C)/(W + C)]s.











