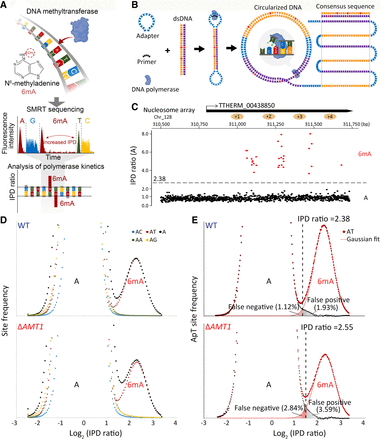
Exclusive methylation at the ApT dinucleotide in Tetrahymena. (A) Overview of 6mA detection by SMRT sequencing. (B) A schematic diagram for SMRT CCS. (C) IPD ratios (IPDr) for all A sites in a typical SMRT CCS read mapped to the Tetrahymena MAC reference genome. The IPDr threshold was set at 2.38, separating 6mA from unmodified A. Note the localization of 6mA clusters in linker DNA between the canonical nucleosome array within the gene body. (D) IPDr distributions (log2) of all A sites in wild-type (WT) (top) and ΔAMT1 cells (bottom). Also plotted were distributions for A sites at the ApA, ApC, ApG, and ApT dinucleotide, respectively. (E) Deconvolution of the 6mA peak and the unmodified A peak for IPDr distributions (log2) at the ApT dinucleotide. Note the low false positive and false negative rates of 6mA calling in WT (top) and ΔAMT1 cells (bottom).











