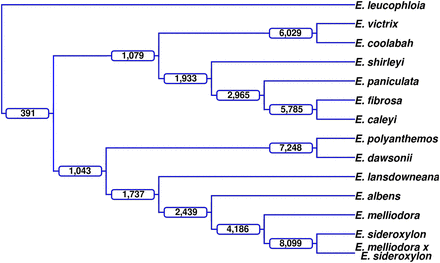
Lineage-conserved rearrangements. Using an outgroup genome, Eucalyptus leucophloia, rearrangements were identified that were shared among members of a lineage, that is, rearrangements with the same start and end points within the outgroup genome. At each branch in the dendrogram, the number of rearrangements shared by all taxa within that clade are labeled; for example, E. victrix and E. coolabah share 6029 rearrangements.











