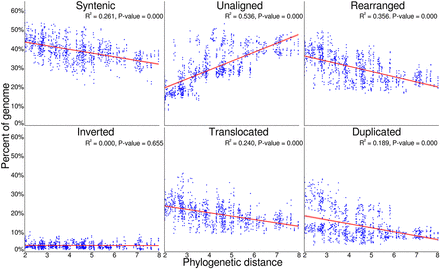
Pairwise genome conservation and loss, as phylogenetic distance increases. The proportion of both Eucalyptus genomes with an alignment pair that was identified as syntenic, rearranged, or unaligned, plotted against the phylogenetic distance of the two genomes. The unaligned proportion is the species-specific fraction of the genome between genome pairs, resulting from either an insertion, deletion, differential inheritance, or sequence divergence. When combined, the proportion of sequence that is syntenic, unaligned, and rearranged equals 100% for each genome within an alignment pair. The rearranged fraction is further broken down into inverted, translocated, and duplicated regions. Phylogenetic distance was calculated as the sum of branch lengths between each genome pair within phylogeny. P-value tests if the slope of the regression line is nonzero.











