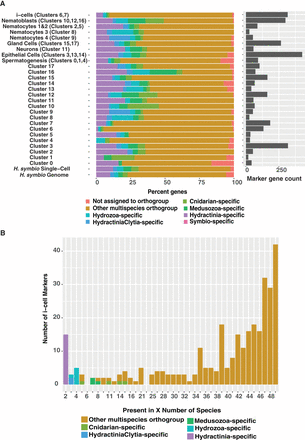
Results from the lineage-specificity analysis using OrthoFinder results and the UMAP cluster marker genes. (A) Stacked bar chart showing the percentage of H. symbiolongicarpus single-cell atlas cluster markers shared among animal phyla. The bottom legend shows eight different categories, dividing the markers into different groups depending on how the orthologs are shared among the species. The “not assigned to orthogroup” category represents markers that could not be placed into an orthogroup. The other categories are markers that have at least one homolog between H. symbiolongicarpus and that category, except for the “symbio-specific” category, which represents markers that fell into orthogroups containing only H. symbiolongicarpus genes. For example, hypothetical marker gene A from H. symbiolongicarpus would be an “other multispecies orthogroup” marker if it was found in H. symbiolongicarpus and at least one animal outside of cnidaria, but it would be a “Cnidarian-specific” marker if it was found in H. symbiolongicarpus and at least one cnidarian outside the Medusozoa. Stacked bars represent the seven major cell types split into nine groups, followed by all individual clusters and, finally, the total genes expressed in the Hydractinia single-cell data set (16,069 genes) and total genes predicted from the Hydractinia genome (22,022 genes). The marker gene count bars on the right indicate how many markers are present in each major cell type and cluster. (B) Histogram dividing the 317 orthogroup-assigned i-cell (clusters C6 and C7) markers by how many are shared by a given number of species. Legend is the same as for panel A, but the following categories are excluded from this chart: unassigned genes (two genes) and H. symbiolongicarpus-specific genes (none).











