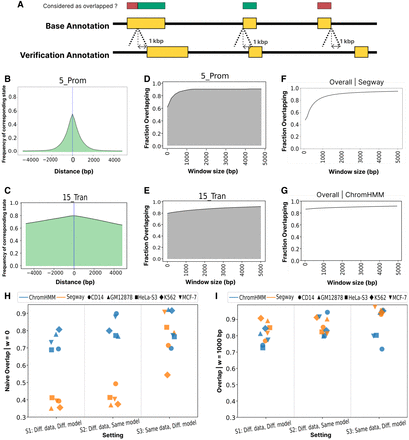
Spatial misalignment of corresponding chromatin states. (A) Schematic depicting the types of state misalignment and the influence of the tolerance window w. The horizontal axis depicts a genomic window. Yellow rectangles indicate a base state and its corresponding verification state. Red/green rectangles indicate whether the given position in the base annotation would be counted as overlapping the verification replicate. (B) Given that a position g is annotated as base state 5_Prom(B), the probability that a nearby position is annotated as the corresponding verification state 13_Prom_fla(V) as a function of distance from g. (C) Same as B, but for 15_Trans(B). (D) Given that a position g is annotated with base state 5_Prom(B), the probability (vertical axis) that the corresponding label occurs within a window g ± w as a function of w (horizontal axis). (E) Same as D, but for 15_Trans. (F,G) The overall overlap of the base annotation from Segway and ChromHMM, respectively, as a function of w (GM12878, setting 1). (H) Naive overlap across two SAGA models, five cell types, and three settings. Color denotes the SAGA model, and shape represents cell type. (I) Same as H, but allowing for a spatial tolerance window of w = 1000.











