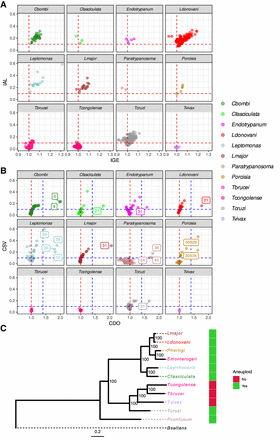
Aneuploidy is ancestral and variable in trypanosomatids. (A) Evaluation of CCNV among trypanosomatid isolates. Each dot corresponds to an isolate. The x- and y-axes represent, respectively, IGE and IAL for each isolate. The vertical red dashed line in the x-value of one represents the expected copy number for euploid samples, with one copy for each haploid genome copy. The horizontal red dashed line corresponds to the IAL of 0.1, representing isolates with relevant CCNV variability. (B) Evaluation of CCNV in each chromosome in the population. Each dot corresponds to a different chromosome. The x- and y-axes represent, respectively, the CDO and CSV of each chromosome across each population. Chromosomes with CDO > 1.4 are identified by their numbers, representing chromosomes with consistent extra copies. The horizontal red dotted line represents the expected CDO value for euploid chromosomes. The horizontal and vertical blue dotted lines correspond, respectively, to CSV 0.1 and CDO 1.4. (C) Trypanosomatid maximum likelihood phylogeny, highlighting the clades that are aneuploid/mostly euploid. Bodo saltans (Bsaltans) was used as an outgroup to root the tree. Node numbers in the tree correspond to the percentage of bootstrap support. The color strip represents the presence/absence of aneuploidies in a given clade. Heatmaps representative of the CCNV for all isolates can be seen in Supplemental Figure S1.











