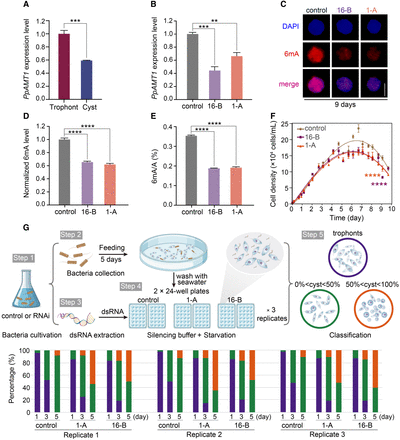
PpAMT1 knockdown (KD) reduces 6mA levels and accelerates encystation. (A) Expression levels of PpAMT1 in trophonts and cysts, calculated by normalized quantitative RT-PCR data. (***) P < 0.001 (Student's t-test). (B) Expression levels of PpAMT1 in control and PpAMT1-KD cells fed for 9 d. Expression levels were calculated by normalized quantitative RT-PCR data. (**) P < 0.01, (***) P < 0.001 (Student's t-test). (C) Immunofluorescence staining of 6mA in control and PpAMT1-KD cells fed for 9 d. Scale bar, 10 µm. (D) Statistical analysis of 6mA immunofluorescence images in C. (****) P < 0.0001 (Student's t-test). (E) Mass spectrometry analysis of 6mA levels in control and PpAMT1-KD cells fed for 9 d. 6mA ratio (6mA/A, %) was defined as the abundance of methylated adenine divided by the total number of adenine. (****) P < 0.0001 (Student's t-test). (F) Growth curves of control and PpAMT1-KD cells. Cell densities were counted using a counting chamber at indicated time points. (****) P < 0.0001 (two-way ANOVA test). (G) Assessment of cyst formation in control and PpAMT1-KD cells. (Top) Workflow of cell feeding, starvation, and RNAi treatment. (Bottom) Statistical results of encystment progression. Cells were fed with E. coli HT115 for 5 d, adding fresh bacteria on a daily basis. After washing with fresh seawater, cells were transferred to two 24-well plates and treated with silencing buffer containing dsRNA daily. The proportion of cysts in individual wells was counted at the indicated time points (n = 48 wells for each replicate) (Supplemental Table S3). The experiment was repeated three times under the same conditions.











