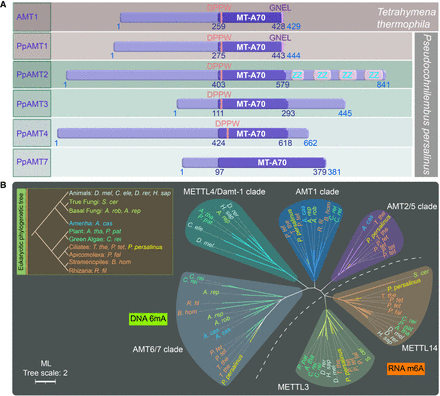
Phylogenetic analysis and domain structure comparison of PpAMT proteins. (A) Domain structures of PpAMT proteins. MT-A70 domains of PpAMT1–PpAMT4 were predicted by CD-Search, whereas that of PpAMT7 was inferred from sequence alignment with PpAMT1–PpAMT4. The structure of AMT1 (MT-A70 family methyltransferase in T. thermophila) is also shown for comparison. The MT-A70 domain, DPPW motif, and the conserved GNEL motif in AMT1 and PpAMT1 are highlighted. (B) Phylogenetic analysis of MT-A70 proteins. The DNA 6mA (clades AMT2/5, AMT1, METTL4/DAMT1, and AMT6/7) and RNA m6A (clades METTL3 and METTL14) methyltransferase candidates are separated by a dotted line. Species are marked by different colors based on their phylogenetic position in the eukaryotic tree. PpAMT1–PpAMT4 and PpAMT7 of P. persalinus are shown in yellow. The scale bar corresponds to two expected amino-acid substitution per site. For details, see Supplemental Table S7 (species full name and NCBI GenBank number).











