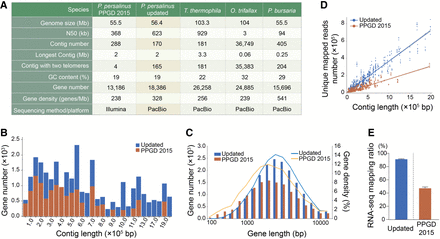
Updated genome assembly of P. persalinus. (A) Statistical details of the new P. persalinus MAC genome assembly and comparison to the PPGD 2015 assembly (Xiong et al. 2015) and three other sequenced ciliate genomes (Lindblad et al. 2019; Sheng et al. 2020; Pan et al. 2023), including Oxytricha trifallax and Paramecium bursaria. (B) Distribution of genes in contigs in the updated genome assembly and PPGD 2015. The x-axis represents the contigs ordered by length; the y-axis, the number of genes. (C) Length distribution of genes in the updated genome assembly and PPGD 2015. The x-axis represents the length of genes; the y-axis, the number/density of genes. Lines represent the gene density; columns, the gene number. (D) RNA-seq mapping variation between the updated genome assembly and PPGD 2015. The x-axis represents the length of contigs; the y-axis, the number of unique reads mapped to the corresponding contigs. (E) RNA-seq mapping ratio of the two genome assemblies. RNA-seq data of trophonts and cyst were mapped to the updated genome assembly or PPGD 2015, respectively.











