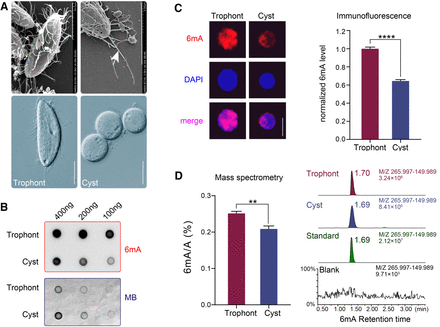
6mA levels are reduced in P. persalinus cysts. (A) Photomicrographs of trophonts and cysts of P. persalinus. (Top) Scan electron microscopic images. (Bottom) Phase-contrast images. Note the magnification difference in the top right image. The arrow indicates the caudal cilium. Scale bars, 10 µm. (B) 6mA levels in genomic DNA of trophonts and cysts were detected by dot blot using an anti-6mA antibody (top). Methylene blue hydrate staining was performed to determine the amount of loaded DNA (bottom). (C) Immunofluorescence staining of 6mA (left) and statistical analysis of 6mA signal intensities (right) in trophonts and cysts. Cell images were randomly selected (n = 100) and processed by ZEN 3.0. (****) P < 0.0001 (Student's t-test). Scale bars, 10 µm. (D) Mass spectrometry analysis of 6mA in genomic DNA of trophonts and cysts. The 6mA ratio (6mA/A, %) was defined as the abundance of methylated adenine divided by total adenine. (**) P < 0.01 (Student's t-test). Chromatograms showing the 6mA mass signal in standard, blank (water), and genomic DNA of trophonts and cysts (right).











