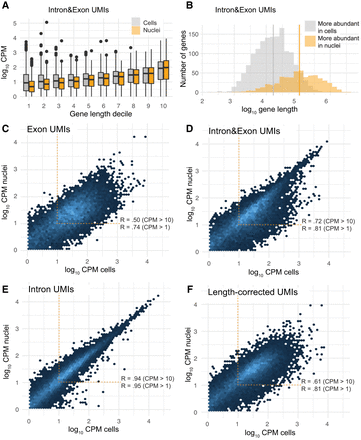
RNA sampling properties explain the effect of bioinformatic processing on assay similarity. (A) Gene length bias is evident in averaged gene abundances in both L5 IT cells and nuclei under the Intron&Exon quantification but is stronger in nuclei. Pearson's correlations between log10 (mean CPM) and gene length are R = 0.51 in nuclei and R = 0.29 in cells. (B) The gene length distribution of genes that are significantly more abundant (fold change >1.5, adjusted P-value < 0.05) either in L5 IT cells (gray) or in nuclei (orange). Mean log10 gene lengths are 5.0 versus 4.2, P < 2.2 × 10−16 (t-test on log10 lengths). (C) Hexbin plot showing the correlation of Exon abundances between L5 IT cells and nuclei. Pearson's correlations are computed on log10 (mean CPM across all cells or nuclei) for genes above one or 10 mean CPM in both assays. (D) Intron&Exon abundances are more strongly correlated and show fewer total differences. (E) The correlation of Intron abundances is very high, consistent with pre-mRNA localization within the nucleus, which is within the cell. (F) Length-corrected abundances are no better correlated than the baseline result. Total differences increase, consistent with the worsened correlation among more highly expressed genes. The length-correction method depresses Intron counts, which indirectly amplifies the prominence of Exon.











