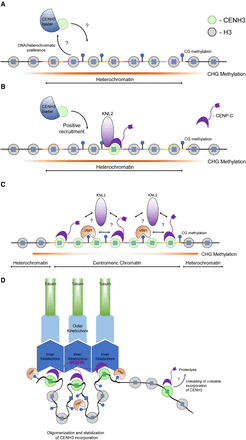
Model for CENH3 loading and centromere homeostasis in Arabidopsis. (A) CENH3 (green circle) is loaded into nucleosomes in a region of CEN178 satellite repeats. The individual repeats are indicated by varying colors. Loading is mediated by a histone chaperone, for example, NASPSIM3 (Le Goff et al. 2020). The region shown is heterochromatic and occupied by nucleosomes containing canonical histone 3 (H3; gray circles). These nucleosomes are modified by H3K9me2, and the associated DNA is methylated in CG, CHG, and CHH sequence contexts. It is possible that heterochromatin marks recruit the CENH3 loading complex. (B) Once CENH3 has been loaded, it assembles kinetochore proteins, including CENP-C and KNL2 (purple) (Ogura et al. 2004; Sandmann et al. 2017; Le Goff et al. 2020). Interactions between the CENH3 loader complex and kinetochore factors may create feed-forward recruitment of CENH3. (C) As the density of CENH3-containing nucleosomes increases, the region undergoes stabilization and compaction of the centromeric chromatin, potentially facilitated by oligomerization of inner kinetochore proteins, for example, CENP-C or KNL2 (Hara et al. 2023; Sissoko et al. 2024). VIM1 (orange) is known to bind and maintain methylation at CG sites and contributes to centromere “strength,” potentially via promotion of kinetochore multimerization. Once a high density of CENH3 nucleosomes is acquired, the region becomes centromeric chromatin, which in Arabidopsis shows reduced CHG context DNA methylation (Naish et al. 2021). (D) Stable loading and maintenance of CENH3 nucleosomes allow recruitment of inner and outer kinetochore complexes (blue) that attach to spindle microtubules (green). We illustrate looping of chromatin via kinetochore multimerization such that CENH3 nucleosomes are gathered in space. Outside of this region, CENH3 and kinetochore proteins are unstable owing to the action of removal pathways, potentially including SUMO or ubiquitin-mediated proteolysis.











