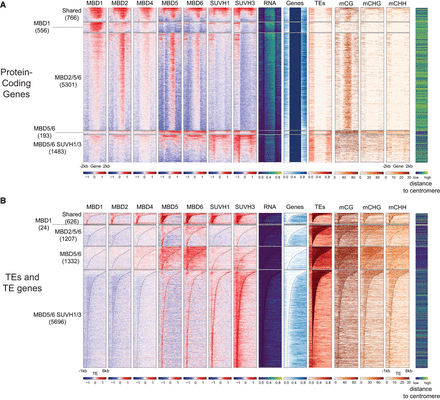
mC readers differ in their binding preferences across genes and TEs. (A) Heatmaps of ChIP-seq signal enrichment of mC readers in protein-coding genes (TAIR10). For each cluster from Figure 2, all unique protein-coding genes intersecting at least one peak were selected, and within each cluster, they were sorted by descending means of the total read number in all bins of the region. For each locus, the methylation level in each context and the RNA expression level (normalized by RPKM) were calculated from the Col-0 WT seedlings sample. For the methylation data, bins with no WGBS coverage are displayed in gray. ChIP-seq signal enrichments were calculated by comparing each ChIP sample to its corresponding input DNA control, scaled in log2 Fold Change (FC) from −1.5 to +1.5. The number of loci in each cluster is reported on the left-hand side of the heatmaps. The presence of an annotated gene or TE in each bin is shown in blue and red, respectively (0 if absent, 1 if present). Genes were scaled to 2 kb, with 1 kb upstream of the TSS and downstream from the TES, plotted in 100 bp windows for DNA methylation tracks and 20 bp windows for the others. (B) Heatmaps of ChIP-seq signal enrichment of mC readers in TEs (TAIR10). For each cluster from Figure 2, all unique TEs and TE genes intersecting at least one of the peaks were selected, and within each cluster, they were sorted by descending means of the total read number in all bins of the region. Heatmap details as in A, except that TEs were aligned on their 5′ end, plotting 1 kb upstream and 6 kb downstream.











