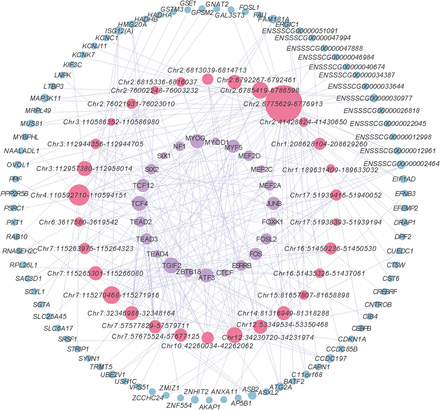
Potential trait-affecting regulatory network mediated by candidate critical enhancers. Lavender nodes represent TFs, pink nodes represent candidate critical enhancers, and light blue nodes represent target genes. The size of each node is proportional to its degree, signifying that larger nodes have more connections. Regulatory interactions flow from TFs to enhancers to target genes. The initial network was constructed by integrating TF motif scanning, TF-target gene coexpression, and eRNA-target gene coexpression. The final network was refined through a series of filtering steps: SE identification, eRNA identification, GWAS hit association, regulatory SNP identification, and TAD filtering.











