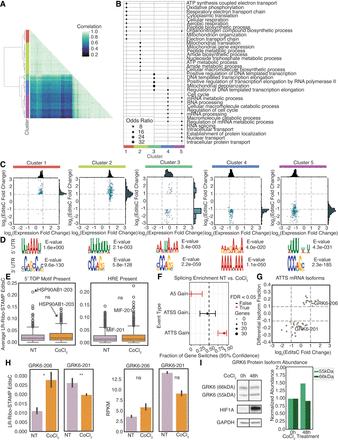
LR-Ribo-STAMP can profile changes in translation at the mRNA isoform level for cells in disease state. (A) Hierarchical clustering of mRNA isoforms based on LR-Ribo-STAMP EditsC and RPKM metrics. The color bar indicates correlations, and the annotations indicate clusters. (B) Gene Ontology enrichment analysis by cluster. (C) Cluster-specific changes in isoform translation (log2(EditsC fold change)) versus change in expression (log2(expression fold change)) following hypoxia. (D) The top enriched motif found in 5′-UTR (top) and 3′-UTR (bottom) sequences for isoforms in each cluster. (E) Translation in normoxia and hypoxia conditions for mRNA isoforms containing 5′TOP motifs (left) and HSEs (right). (F) Splicing event enrichment following hypoxia. (G) Differences in isoform fraction usage (DIF) versus change in translation following hypoxia. (H) LR-Ribo-STAMP of EditsC (left) and isoform expression (RPKM; right) of GRK6-206 (ENST00000507633) and GRK6-201 (ENST00000355472) mRNA isoforms. (I) Western blot of the protein isoforms that result from GRK6-206 (55 kDa) and GRK6-201 (66 kDa), at the 0 h and 48 h of hypoxia. Values are normalized against the 0 h timepoint. Significance: (***) P ≤ 0.001, (**) P ≤ 0.01, (*) P ≤ 0.05.











