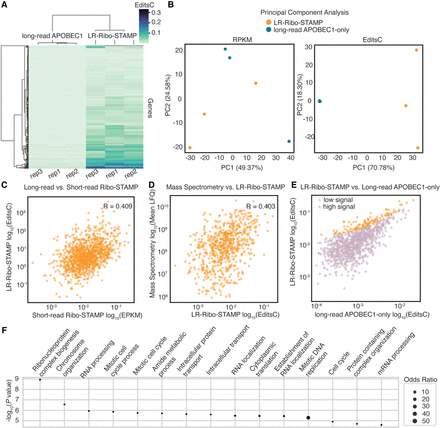
LR-Ribo-STAMP profiles translation at the gene level with good concordance with short-read data. (A) Signal detection (EditsC) across replicates of LR-Ribo-STAMP and long-read APOBEC1-only samples at the gene level. (B) PCA plot showing clustering of LR-Ribo-STAMP (gold) and long-read APOBEC1-only (green) samples based on gene-level RPKM (gene expression; left) and EditsC (editing; right) metrics. (C) Spearman's rank correlation of LR-Ribo-STAMP EditsC computed from long-read sequencing data and Ribo-STAMP EPKM computed from short-read sequencing data. (D) Spearman's rank correlation of LR-Ribo-STAMP and mass spectrometry data collected from unperturbed HEK293T cells. (E) Results from a linear model built using gene-level EditsC from LR-Ribo-STAMP and long-read APOBEC1-only. The results delineate genes with high signal (orange) over the background (pink). (F) Gene Ontology enrichment of genes with high signal as designated by the linear model, ranked by −log10(P-value).











