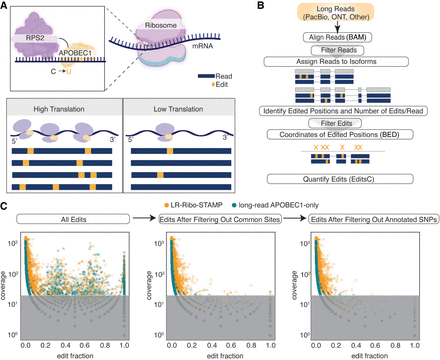
Experimental and computational methods for long-read Ribo-STAMP (LR-Ribo-STAMP). (A) Overview of the LR-Ribo-STAMP experimental system. RPS2 is fused to APOBEC1 to induce cytosine-to-uracil nucleotide edits proximal to ribosome–RNA interaction sites. More edits indicate higher translation, and fewer edits indicate lower translation. (B) Overview of the LR-Ribo-STAMP computational pipeline. The input is unaligned long reads, which undergo alignment, read filtering, edit detection, edit filtering, and edit quantification. Edited sites are output as a BED file. (C) Edit filtering. An outline of edit filtering and delineation of LR-Ribo-STAMP (gold) from long-read APOBEC1-only (green) signal through filtering common sites and annotated SNPs represented as the relationship between edit fraction (edited reads/total number of reads) versus coverage at an edited site. Edited sites in the gray portion indicate sites having fewer than 20 reads filtered out.











