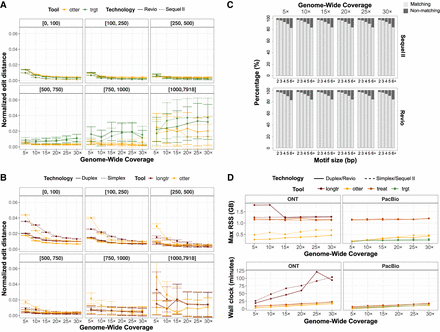
Benchmarking between TREAT/otter, TRGT, and LongTR. (A) Genotyping accuracy of otter and TRGT on PacBio Sequel II and Revio data, stratified by TR size, and sequencing depth. (B) Genotyping accuracy of otter and LongTR on ONT Simplex and Duplex data, stratified by TR size and sequencing depth. (C) Motif identification accuracy of TREAT and TRGT on PacBio Sequel II and Revio data, showing the overlap of matching motifs, stratified by motif size and sequencing depth. (D) Memory usage and running time of otter, TREAT, TRGT, and LongTR, across technologies and sequencing coverages.











