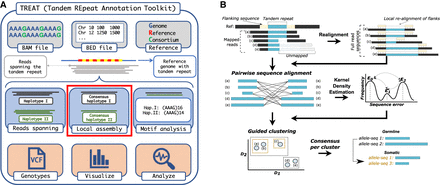
Figure 1.
Schematic workflow of TREAT and otter. (A) TREAT workflow, highlighting the required inputs, the main features, and the main outputs of the tool. The red box highlights the main genotyping engine based on otter. (B) The main algorithmic steps of otter, a novel targeted local assembler for long-read sequencing data.











