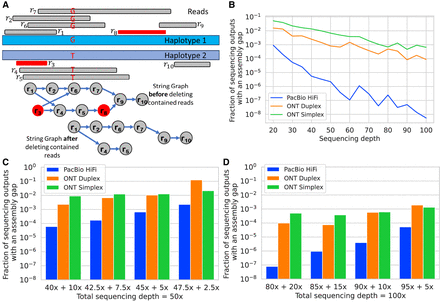
Assembly gaps and their occurrence frequency. (A) An example of a sequencing output where an assembly gap occurs in the string graph due to contained read deletion. Read r3 is contained in read r1. Read r8 is contained in read r7. Accordingly, the string graph representation excludes reads r3 and r8. Read r3 is redundant; its deletion simplifies the graph. However, removing read r8 breaks the connectivity between reads r5 and r9, which was necessary to spell the second haplotype. (B) Fraction of sequencing outputs containing an assembly gap. We measured the fractions using the read-length distributions corresponding to three sequencing technologies (PacBio HiFi, ONT Duplex, ONT Simplex) and using different sequencing depths. Here, we used equal sequencing depths on both haplotypes. (C,D) Fraction of sequencing outputs containing an assembly gap when the sequencing depths across the two haplotypes are uneven. This scenario models somatic mutation in DNA with variant allele frequency below 0.5. In (C), the total sequencing depth for both haplotypes is 50×. In (D), the total sequencing depth is 100×.











