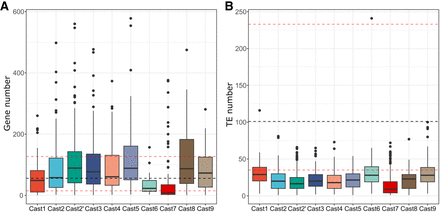
Distribution of genes and TEs in the vicinity of Cast arrays. (A) The number of genes was calculated based on the distribution of gene content in the assembled genome, with the black line representing the median number of genes in regions of same size (100 kb windows, 5 kb sliding frame) and the red dotted lines representing the first and third quartiles (Q1 and Q3) of the genome-wide distribution. Results of the Kolmogorov–Smirnov tests are represented in Supplemental Table S12. (B) The number of TEs was calculated using the same window size as for the genes, with black and red lines representing the same values. All Cast arrays had significantly (Kolmogorov–Smirnov P < 0.01) fewer TEs in their vicinity compared with the genome-wide distribution, highlighting the fact that they do not overlap with TE presence.











