Figure 1.
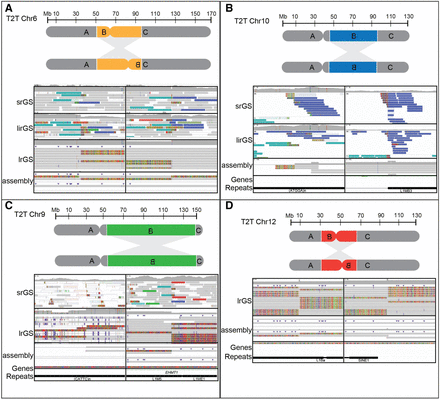
Reference genome-dependent detection of inversions analyzed by srGS and lrGS. (A) An inversion 6 (P4855_501) visible in srGS, linked read genome sequencing (lirGS), and lrGS using GRCh38. (B) An inversion 10 (P4855_106) visible in srGS and lirGS data using T2T-CHM13. (C) An inversion 9 (BH16643-1) only visible by lrGS de novo assembly using T2T-CHM13. (D) An inversion 12 (RD_P541) within a 8 kbp DRR.











