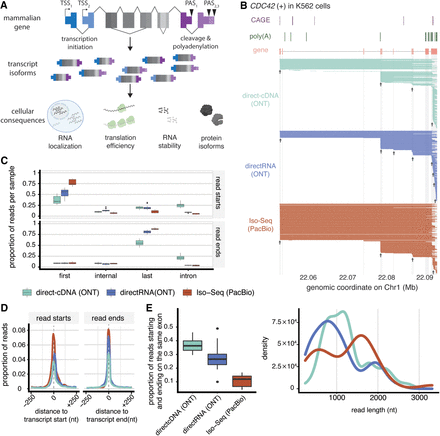
LRS reads show large variability in start and end coordinates. (A) Mammalian genes can have multiple alternative transcript starts (TSSs) and ends (PASs), leading to different isoforms expressed from the same gene with variable cellular consequences. (B) Representative example of long-read RNA sequencing reads for CDC42 in K562 cells. To aid visualization, 280 randomly sampled reads from each sequencing technology are shown. At the top are annotated features of CDC42, including CAGE peaks (violet), poly(A) peaks (dark green), and annotated exons (orange). At the bottom are read coverage plots, in which each horizontal line represents the span between the first and last coordinates of a read, for the ONT direct-cDNA (teal) (Chen et al. 2021), ONT directRNA (blue) (Chen et al. 2021), and PacBio Iso-Seq (red) (Luo et al. 2020) data sets (Supplemental Methods; Supplemental Table S1). Arrows mark TSS clusters. The bottom panel shows the distribution of read length across sequencing technologies for CDC42. (C) Distributions of the proportion of reads that start (top) or end (bottom) in HITindex-classified (Fiszbein et al. 2022) first, internal, or last exons or introns across from three LRS technologies, using data from the A549, Hct116, HepG2, K562, and MCF-7 cell lines. (D) The distribution of read starts and ends around annotated transcription start (left) or end (right) sites across three LRS technologies. The y-axis represents the proportion of reads per sample, calculated using a sliding window of 0.01 kb around the feature. (E) Distributions of the proportion of single-exon reads that start and end within the same exonic feature across three LRS technologies. Note that B–E use published data sets, and the metrics presented here may change when estimated with other data sets or protocols.











