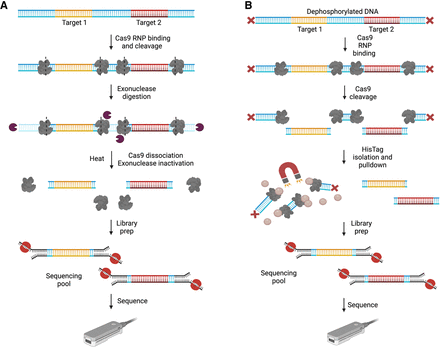
Approaches with background reduction. (A) NE and CaBagE. DNA is cleaved upstream and downstream from the ROI (dashed lines represent cut sites) using target-specific crRNPs. Immediately after Cas9 cleavage, 5′ and 3′ exonucleases are used to digest background DNA while target ends are protected by the bound Cas9. Heat incubation is used to dissociate Cas9 from the targets and inactivate the exonucleases before sequencing adapter ligation. The prepared library is loaded on nanopore flowcells and sequenced. (B) Affinity-based Cas9-mediated enrichment (ACME). DNA is dephosphorylated to prevent sequencing adapter ligation and Cas9 RNPs with guides specific to the ROI are used to cleave the DNA upstream and downstream from the target(s). After cleavage, Cas9 remains bound to the nontarget side of the cut sites (PAM-distal end). The Cas9 enzyme has a C-terminal 6 Histidine Tag. HisTag Dynabeads are used to pull down Cas9 and the nontarget fragments bound to it from the sequencing pool. Adapters are then ligated to the exposed phosphate groups at the ends of the target strand(s). The prepared library is sequenced on ONT flowcells. (Created with BioRender; https://www.biorender.com/)











