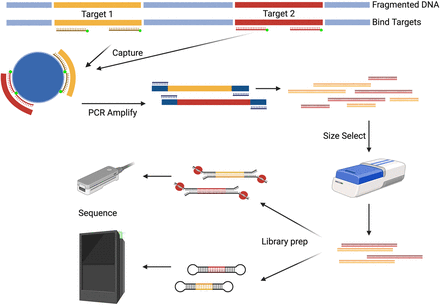
Hybridization-based capture. Biotinylated DNA or RNA guides are designed to be complementary to the ROI. The DNA is fragmented to ∼10 kb and amplified if more mass is needed. Next, the probes bind to the denatured DNA. The probe–ROI complex undergoes a bead-based pulldown to separate the target regions from the rest of the genome. The enriched fragments are amplified and size-selected to maintain the target length. The amplicons are then prepared for long-read sequencing. (Created with BioRender; https://www.biorender.com/)











