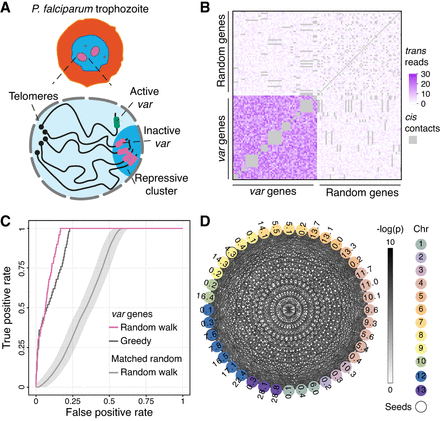
Trans-C identifies the var genes cluster in Plasmodium falciparum. (A) Schematic of P. falciparum in the trophozoite stage of its red blood cell life cycle, with a zoomed-in view of the nucleus highlighting its Rabl-like structure and the clustering of the var genes in a repressive heterochromatic cluster. (B) Contact heat map comparing trans contact counts among all 60 var genes versus 60 randomly selected 10 kb bins. Cis contacts are grayed out. (C) Performance evaluation of trans-C-mediated identification of var gene clustering. We plot the receiver operating characteristic (ROC) curve for the trophozoite life stage of P. falciparum. The var genes are uncovered by the random walk algorithm of trans-C with high area under the ROC curve (AUROC; 0.94). The cumulative distribution is statistically significant (P = 3 × 10−165) from a null model of 1000 random walks performed from seeds selected randomly but with an equal or greater collective interaction strength (matched random; the line reports the average and the shaded area the 95% confidence interval). We also report the performance of a simpler greedy heuristic. (D) Visualization of the var gene–associated trans clique identified by trans-C in P. falciparum trophozoite. Nodes are color-coded by chromosome and sequentially numbered based on their relative position along each chromosome (expressed in megabases). Edges are color-coded based on trans interaction significance (cis contacts are not plotted). The seed loci for the random walk are indicated by solid black lines around the nodes.











