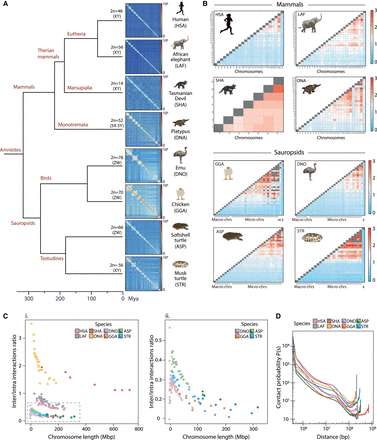
Patterns of chromosome folding across vertebrates. (A) Whole-genome Hi-C contact maps for human (Homo sapiens [HSA], Rao et al. 2014), African Elephant (Loxodonta africana [LAF], Álvarez-González et al. 2022a), Tasmanian Devil (Sarcophilus harrisii [SHA], Álvarez-González et al. 2022a), Platypus (Ornithorhynchus anatinus [ONA], Zhou et al. 2021), Emu (Dromaius novaehollandiae [DNO], Liu et al. 2021), chicken (Gallus gallus [GGA], Fishman et al. 2019), Spiny Softshell Turtle (Apalone spinifera [ASP], this study), and Northern Giant Musk Turtle (Staurotypus triporcatus [STR], this study]. (B) Heat maps representing interchromosomal mean interactions per chromosome for human, African Elephant, Tasmanian Devil, Platypus, chicken, Emu, Spiny Softshell Turtle, and Northern Giant Musk Turtle. (Ci) Inter-/intrachromosomal interactions as a function of chromosome length (in Mbp) in the target mammals and sauropsids compared in this study. In the case of turtles and birds, color intensity differentiates between macro (darker color) and micro (lighter color) chromosomes. (Cii) Enlarged view of the inter-/intrachromosomal interactions analyzed for birds and turtles (demarcated by the dashed line in Ci). (D) Chromosome-specific contact probability P(s) as a function of genomic distance in all species analyzed.











