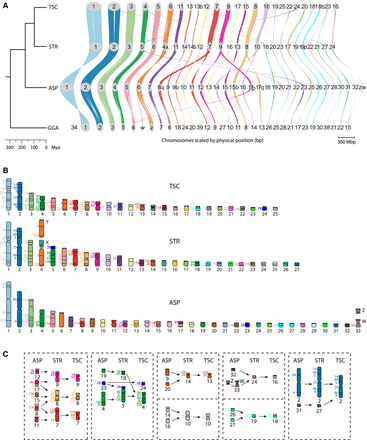
Genome conservation and chromosomal rearrangements. (A) Genespace plot depicting chromosomal syntenies (of sequences >1 Mb) and rearrangements (of >5 genes) between turtles and chicken. Chromosomes are scaled by physical size (bp). (TSC) Trachemys scripta elegans assembly from Simison et al. (2020) with corrected chromosomal nomenclature as per (Lee et al. 2020), (STR) Staurotypus triporcatus (this study), (ASP) Apalone spinifera (this study), (GGA) Gallus gallus (Warren et al. 2023). Suffixes “p” and “q” denote a fragment of the short- and long-arm of a chromosome, respectively, whereas “b” denotes a fragment of an acrocentric chromosome or of the same chromosome arm. (B) Ideograms depicting BAC positions and chromosomal homology across species (Badenhorst et al. 2015). (C) Major chromosomal fusions and fissions identified in this study between Apalone, Staurotypus, and Trachemys (chromosome nomenclature and homology as in B).











