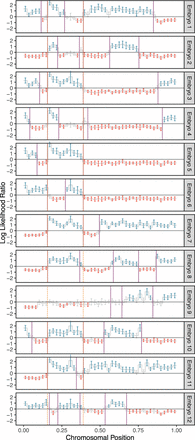
A representative example of crossover discovery based on haplotype matching among sibling IVF embryos. In this example, Chromosome 10 of each test embryo is compared with a single sibling reference embryo with monosomy of Chromosome 10. Evidence of haplotype nonmatching is indicated by positive log-likelihood ratios, whereas evidence of haplotype matching is indicated by negative log-likelihood ratios. Crossovers are identified as transitions from positive (blue) to negative (red) log-likelihood ratios or vice versa. Each point corresponds to a bin size of 3 Mbp and consists of a varying number of genomic windows. Error bars denote 95% confidence intervals. Crossovers attributed to the monosomic chromosome are indicated with dashed orange lines (defined as those observed in more than half of test samples), whereas crossovers attributed to the test samples themselves are indicated with purple lines.











