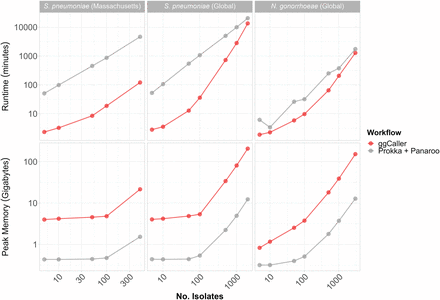
Computational benchmarking of ggCaller against workflow of Prokka + Panaroo. Tools were run using 16 threads, comparing runtime (top) and peak memory (bottom), with an increasing number of randomly sampled genomes. Horizontal panels describe data set S. pneumoniae (Massachusetts), data set from Croucher et al. (2015); S. pneumoniae (global), data set from Gladstone et al. (2019); and N. gonorrhoeae (global), data set from Blackwell et al. (2021). For consistency, the same gene annotation databases were provided to both Prokka and ggCaller for each data set.











