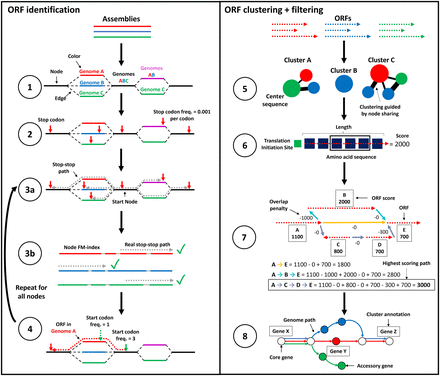
ggCaller workflow. ggCaller can be split into two sections: ORF identification (steps 1–4) and ORF clustering + filtering (steps 5–8). (1) DBG is generated from assemblies by Bifrost. (2) All stop codons are identified, and stop frequency is calculated (total number of stop codons in DBG / total number of codons in DBG). (3a) Starting at an initial node containing a stop codon, a depth first search (DFS) is used to pair all stop codons in the start node with a downstream stop codon in the same reading frame. (3b) During DFS, paths are compared to an FM-index to remove incorrect paths. (4) ORFs are defined by identifying start codons scored based on translation initiation site sequence, genome coverage (given by number of colors shared in node), and frequency of this start being chosen in other potential orthologs. Steps 3 and 4 are repeated for all nodes containing a stop codon. (5) ORFs are clustered into COGs, using node-sharing to reduce search space. (6) Balrog is used to generate an average per-residue score using only the center sequence of each COG. This average per-residue is used to score each ORF in the center sequence's respective cluster. (7) Highest scoring tiling path calculated for overlapping genes within the DBG using the Bellman–Ford algorithm (Bellman 1958; Ford and Fulkerson 1962), producing a “true” gene-call set. (8) Gene-calls and synteny information are used to build a gene graph. A modified version of Panaroo is used to remove poorly supported gene-calls, annotate clusters, and recall missed genes/pseudogenes.











