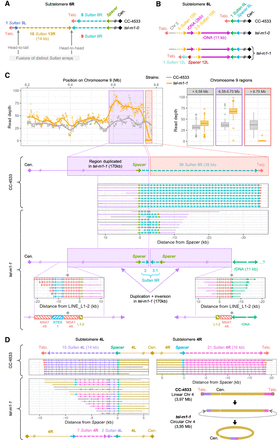
Complex genome rearrangements at subtelomeres. (A) Schematic representation of subtelomere 6R in which, in tel-m1-0, 2 additional Sultan arrays were fused to the initial Sultan array in inverted orientation. (B) Schematic representation of complex rearrangements at subtelomere 6L in tel-m1-0 and tel-m1-1, including Sultan elements from different subtelomeres and rDNA sequences. In tel-m1-1, two subpopulations of reads reveal two distinct structures stemming from the initial rearrangement found in tel-m1-0. (C) Signature of a BFB event at subtelomere 9R in tel-m1-1. (Top) Duplication of the last 170 kb of the chromosome end. Read depth is computed on the indicated regions of Chromosome 9. (Middle) Individual reads supporting the loss of telomeres and 36 Sultan elements, as well as the end-to-end fusion of the 9R sister chromatids. (Bottom) Reads showing the recruitment of >11 kb of rDNA sequences at the new 9R extremity, disrupting an array of MSAT-4B satellite sequences. (D) Representation of reads supporting the fusion of subtelomeres 4L and 4R in tel-m1-1, after complete loss of telomeres and a total of 27 Sultan elements. (Bottom right) Scheme of the inferred circularization of Chromosome 4, also supported by de novo genome assembly (Supplemental Fig. S6E,F). Black segments correspond to sequences that are not aligned to any of the selected queries.











