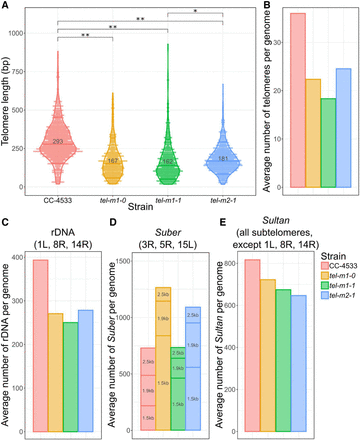
Telomere length distribution and CNVs of subtelomeric elements. (A) Telomere length distribution of terminal telomere sequences detected by TeloReader from the reads data set in CC-4533 and three telomerase mutant samples. Statistically significant difference: (*) P-value = 6.8 × 10−9 and (**) P-value < 2.2 × 10−16, using a nonparametric Kruskal–Wallis test. (B) Number of telomeres in reads normalized by the sequencing depth. (C–E) CNVs for repeated elements normally found at subtelomeres including the rDNA (C), the Suber elements subdivided into three types according to their length (“1.5 kb,” “1.9 kb,” and “2.5 kb”; D), and the Sultan elements (E).











