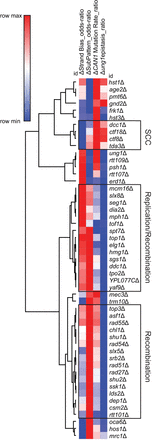
Nonbiased hierarchical clustering of yeast gene deletions that similarly impact A3B-induced mutation. Yeast gene deletion strains with greater than 20 independent A3B-induced CAN1 mutations sequenced were clustered using average linkage based upon (1) how much the gene deletion elevated A3B-induced mutation over that in wild-type yeast, ΔCAN1 Mutation Rate_ratio, which equals the log2(CanR rateyfgΔ/CanR ratewild-type); (2) how the rate of mutation in the yfgΔ ung1Δ codeletion strains compared with the mutation rate in the ung1Δ strains, Δung1epistasis_ratio, which equals the (log2(CanR rateyfgΔ ung1Δ/CanR rateung1Δ); (3) mutational strand bias, ΔStrand Bias_odds-ratio, which equals the log2(substitutions at C/substitutions at G nucleotides); and (4) polymerase usage, ΔSubPattern_odds-ratio, which equals the log2(C-to-T substitutions/C-to-G substitutions). Nodes of genes associated with sister chromatin cohesion (SCC), DNA replication and recombination, and recombination were identified (black boxes).











