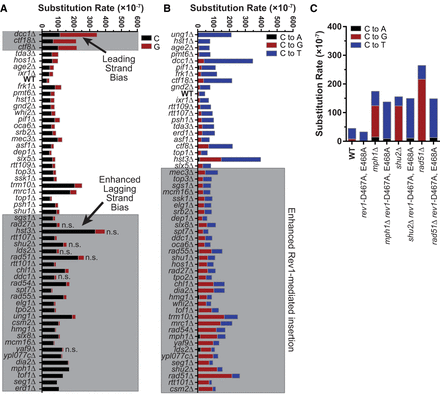
Spectra of A3B-induced mutations in CAN1. The CAN1 gene of independent CanR isolates was amplified and sequenced to determine the spectra of A3B-induced mutations in yeast with specific gene deletions. Note that A3B-induced CAN1 mutants were not sequenced from the aim9Δ, rnr4Δ, ser3Δ, swe1Δ, yap1802Δ, YKL136WΔ, and YHL042WΔ strains. (A) Strand bias of A3B-induced mutations in CAN1 is influenced by specific gene deletions. The rate of C (black bars) or G (red bars) substitutions in each genotype of yeast expressing A3B. Wild-type yeast expressing A3B favors C substitutions over G substitutions, indicating that the lagging strand template is the top DNA strand of CAN1 at its location on ChrV. The ratio of A3B-induced C-to-G substitutions in wild-type yeast was compared to the same ratio for each yeast deletion strain by two-sided Fisher's exact test. Genotypes with P-values < 0.00083 (based on Bonferroni multiple hypothesis testing correction) were considered to have altered strand bias. (B) The rates of C-to-A, C-to-T, and C-to-G substitutions (complementary substitutions were combined) in CAN1 for yeast expressing A3B and containing various gene deletions. The ratio of C-to-T and C-to-G substitutions were compared pairwise between wild-type yeast expressing A3B and each specific gene deletion strain by two-sided Fisher's exact test. Gene deletion strains displaying P-values < 0.00083 (for Bonferroni correction) were deemed to have more A3B-induced abasic sites bypassed via Rev1 catalytic activity. (C) Rates of C-to-A, C-to-T, and C-to-G substitutions (as in B) for comparing WT and recombination-deficient yeast with corresponding strains lacking Rev1 catalytic activity (i.e., rev1-D467A, E468A).











