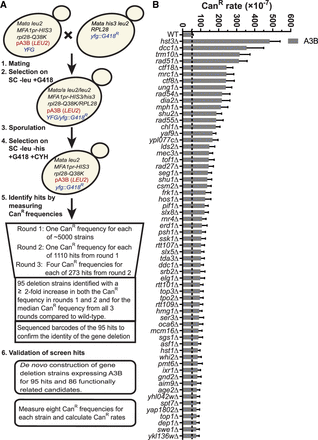
Identification of yeast gene deletions that increase APOBEC3B-induced mutations. (A) Schematic of the yeast screen used to identify gene deletions that elevate A3B-induced mutagenesis. Haploid yeast containing a single-gene deletion and the A3B expression plasmid was obtained by mating MATα yeast carrying the A3B expression plasmid with each MATa strain of the yeast gene deletion library, sporulating the resulting diploid cells, and selecting the desired haploid genotype. The resulting haploid cells were used for three rounds of CanR frequency measurements to identify gene deletions that likely augment A3B-induced mutagenesis. We were unable to generate 21 haploid strains for the screening process that represent potential (unvalidated) synthetic lethal interactions with A3B expression (Supplemental Table S1). To confirm these deletions resulted in increased A3B-induced mutations we recreated each deletion strain expressing A3B de novo and measured CanR mutation rates. (B) CanR rates for 61 yeast gene deletions that elevate A3B-induced mutation more than 1.55-fold over the A3B-induced CanR rate in wild-type yeast (gray dashed line) without overlapping 95% confidence intervals and 1.55-fold higher than additive for spontaneous deletion-dependent CanR and wild type with A3B CanR rates. Error bars indicate 95% confidence intervals. For all mutation rate data, see Supplemental Table S2.











